New method identifies bacteria more easily
Researchers have developed a method that identifies bacteria easily, cheaply and more precisely than before. This can help reduce use of antibiotics.
Far too many antibiotics are used around the world. As a result, bacteria are becoming resistant to these drugs.
Curing bacterial diseases is becoming more difficult than before, because antibiotics are perhaps our foremost weapons in the fight against them.
We have developed a simple tool that can identify all of the genetic material in bacteria.
An important step towards using fewer antibiotics is to find better methods for identifying pathogens, and here is the good news.
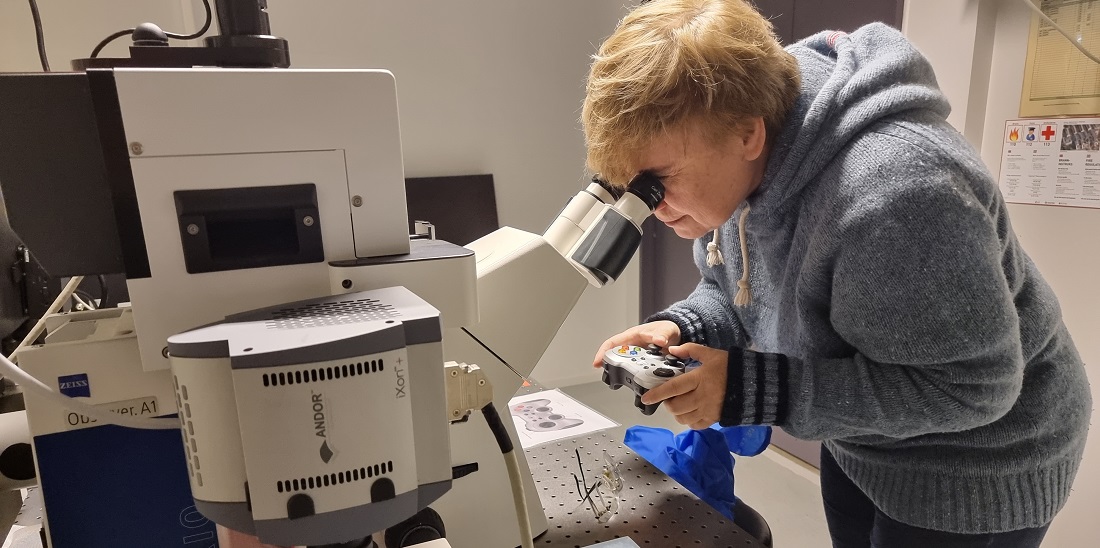
“We have developed a simple tool that can identify all of the genetic material in bacteria. This allows us to find out more quickly what kind of bacteria a sick person or animal is affected by, or what kind of bacteria are found in food or the environment. We can then also decide whether it is necessary to use antibiotics against the bacterium, and if so what kind, so we don’t have to use as much medication,” says Professor Erika Eiser at NTNU’s Department of Physics.
No need to copy genetic material
An international research group is behind the latest findings. The results have been presented in the prestigious Proceedings of the National Academy of Sciences (PNAS) journal. Playing a key role in the work was Peicheng Xu from the Institute of Physics Chinese Academy of Sciences in Beijing, for whom Eiser was previously an academic supervisor.
One reason why the new method is faster is that users do not have to go through a step called ‘gene amplification’. This involves making several copies of the genetic material so it is easier to analyse, but this step can now be skipped.
“We can analyse all of the bacterium’s DNA without gene amplification by using a method previously used in simulations,” says Professor Eiser.
Eiser was part of a research group led by Tine Curk from Johns Hopkins University that developed the theory behind the method, which also works in reality.
“We get excellent results when we apply the theoretical method to real samples,” says Eiser.
The method creates clumps
This paragraph might be a bit difficult to understand, but basically, DNA is made up of rows of so-called nucleotides. The new method enables researchers to find short sequences of the bacteria’s DNA. They do this by seeing how these sequences bind to different variants of DNA that are grafted onto colloids, which are particles dissolved in a liquid.
If you are interested in finding out more, you can read about the process in more detail here. What it means, however, is that researchers can quickly identify the bacteria, because they bind themselves to these colloids in various ways and cause them to clump together.
The bottom line is: you don’t have to analyse so much material. You can skip the step of having to copy them, and this saves time and money.
“Using this method, we saw how as few as five E. coli bacteria caused the colloids to create clusters,” says Professor Eiser.
Still a way to go
All of this is currently in its early stages. Eiser has published a proof-of-principle experiment. This means that there is still a lot of work to be done before it becomes a widely used method.
“The findings can provide us with a reliable method for identifying pathogens in disciplines such as food safety, disease control and environmental monitoring,” says Professor Eiser.
In a world where more and more bacteria are becoming resistant to antibiotics, this is particularly good news.
References: P. Xu, T. Cao, Q. Fan, X. Wang, F. Ye, E. Eiser ‘Whole-Genome Detection using Multivalent DNA-Coated Colloids’ PNAS 120 (37), 2305995120; DOI: https://doi.org/10.1073/pnas.2305995120 (2023)